|
3/25/2023 0 Comments Nokoi peptideshaker 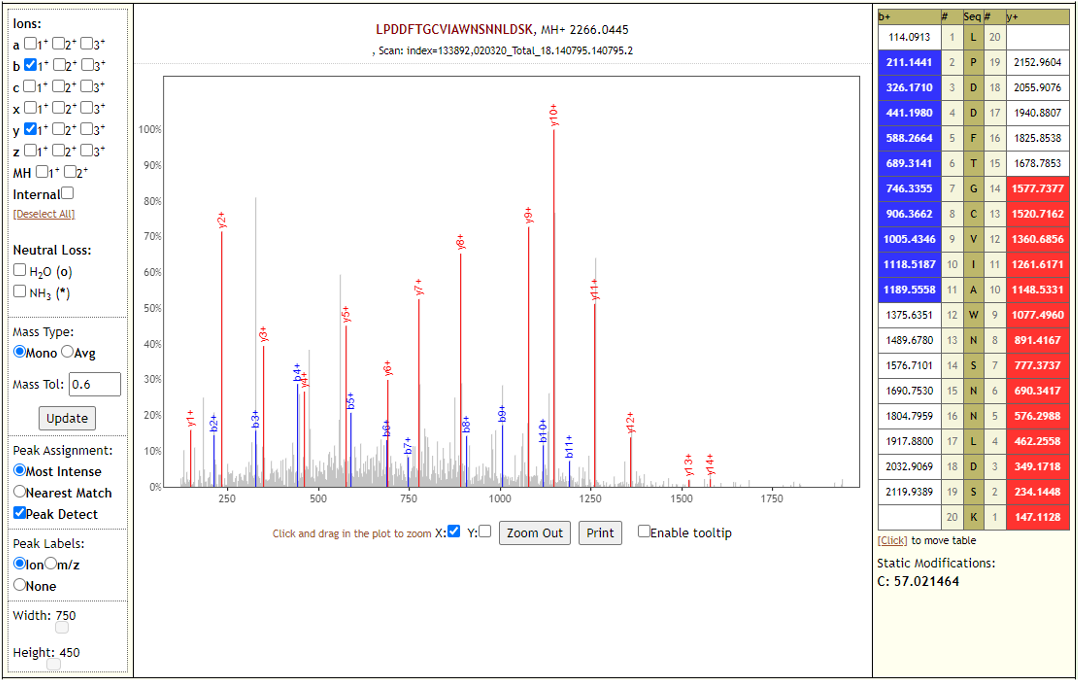
Characterisation of the serum proteome of sheep would therefore be useful to quantify disease in this species. To date, the serum proteome of sheep is largely extrapolated from cattle, which can be inaccurate despite a 97% similarity in protein coding sequences and different promoters driving the expression of specific proteins. However, the well-defined serum proteome of humans permits analysis and interpretation of a range of physiological changes and disease states. There is currently no publicly available reference mass spectrometry-based circulating acellular proteome data for sheep. This peptide spectral data contributes to a protein library that can be used to identify a wide range of proteins in ovine serum. These results demonstrated for the first time the feasibility of characterising the ovine circulating acellular proteome using nanoLC-nanoESI-MS/MS. These data are available via ProteomeXchange with identifier PXD004989. Since Mascot software is considered the industry standard and identified the most proteins, these were analysed using the Protein ANalysis THrough Evolutionary Relationships (PANTHER) classification tool revealing the association of 349 genes with 127 protein pathway hits. ProteinPilot™ and Mascot identified 245 and 379 protein groups (IDs), respectively, and PeptideShaker validated 133 protein IDs from the entire dataset. PeptideShaker (CompOmics, VIB-UGent) searches were used to validate protein identifications from ProteinPilot™ and Mascot. Proteins were identified using ProteinPilot™ (SCIEX) and Mascot (Matrix Science) software based on a minimum of two unmodified highly scoring unique peptides per protein at a false discovery rate (FDR) of 1% software by searching a subset of the Universal Protein Resource Knowledgebase (UniProtKB) database ( ). Serum samples from healthy sheep were subjected to shotgun proteomic analysis using nano liquid chromatography nano electrospray ionisation tandem mass spectrometry (nanoLC-nanoESI-MS/MS) on a quadrupole time-of-flight instrument (TripleTOF® 5600+, SCIEX). The objective of this study was to develop a robust and comprehensive method to characterise the circulating acellular proteome in ovine serum. 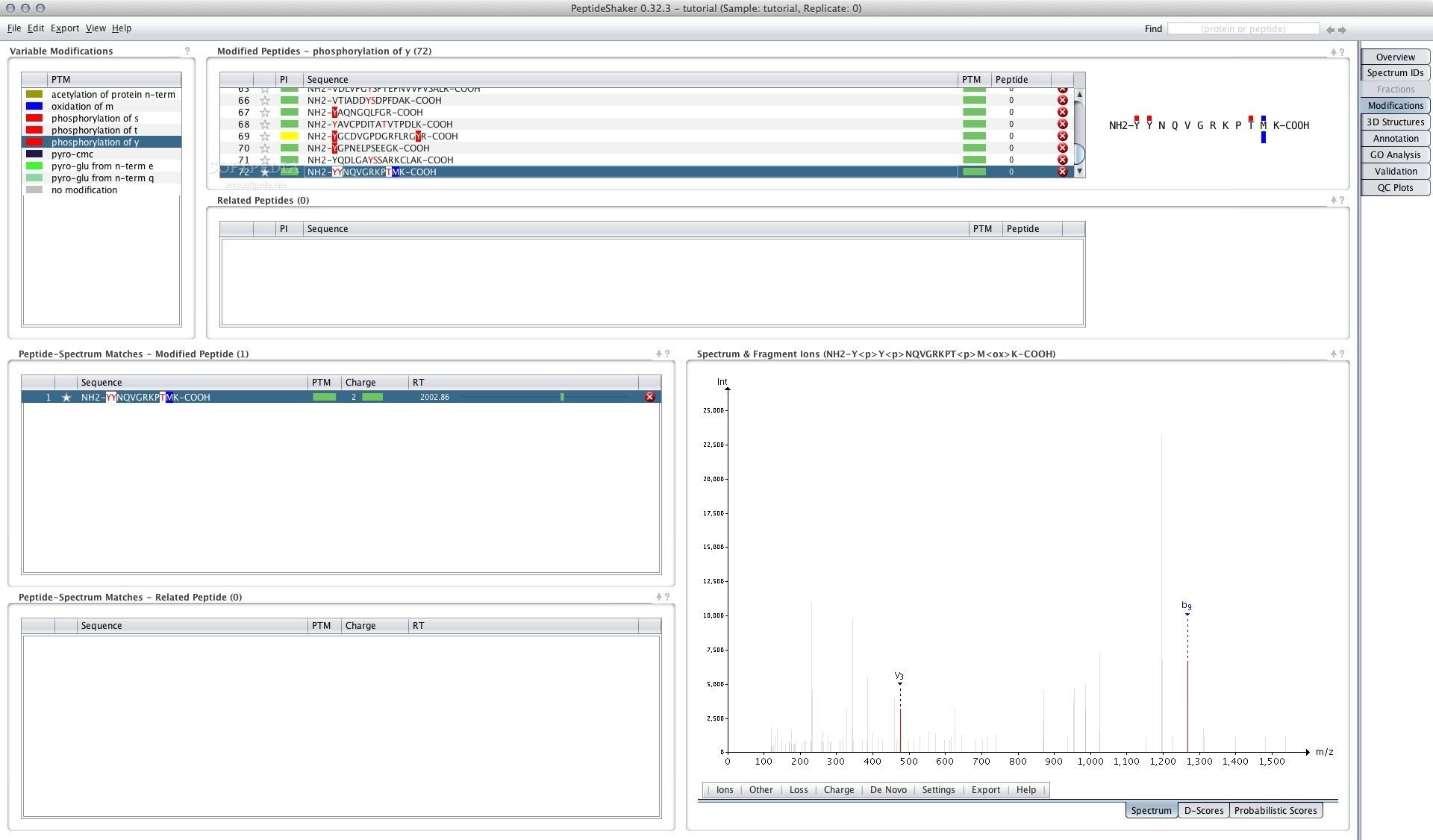 
Unlike humans, there is currently no publicly available reference mass spectrometry-based circulating acellular proteome data for sheep, limiting the analysis and interpretation of a range of physiological changes and disease states. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
